Research
Master's Research in DNA Topology
Abstract
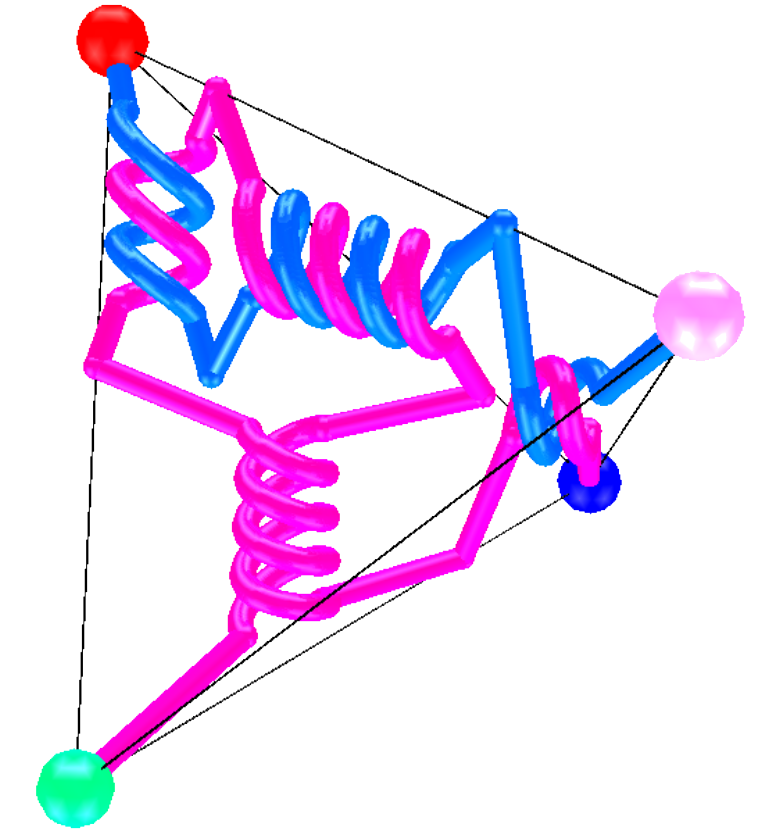
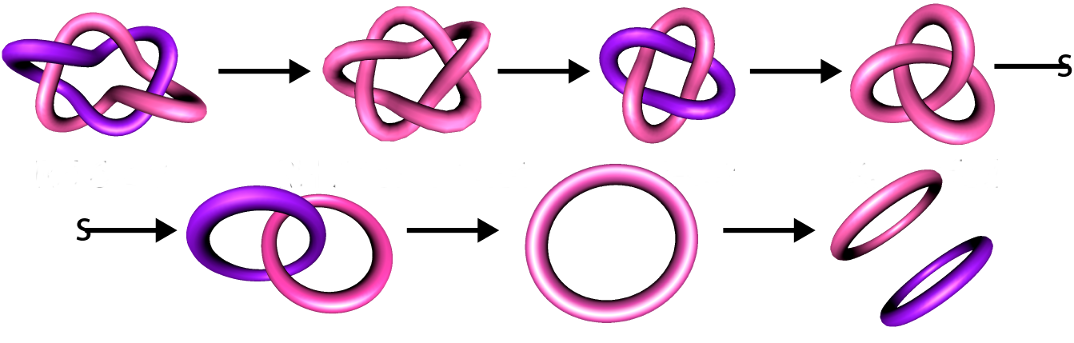
Knotted and linked DNA cause complications during DNA replication and transcription, and therefore simplifying these topological forms is essential. In Escherichia coli, the XerCD-FtsK complex has been found to effectively unlink DNA links produced by replication of the circular chromosome. TangleSolve, a computer implementation of the tangle method of Ernst and Sumners, can be used to compute possible enzymatic mechanisms. In 2005, Vazquez and colleagues showed that the three mechanisms proposed for the action of XerCD on a circular unknotted DNA molecule could be interpreted as different projections of a 3D object. In 2012, Wono extended this idea by entrapping tangles inside a regular tetrahedron. Here we expand TangleSolve, and develop a computer visualization tool called Tangle3D, that automates and extends Wono’s work. Using Tangle3D we define equivalence classes for the projections of tetrahedral tangles and explore an application to DNA unlinking by Xer recombination. Moreno Master's Thesis.
Site-Specific Recombination

XerCD-FtsK Unlinking Steps